Abstract
We have performed an entire-population-based survey of the epidemiology and penetrance of Leber hereditary optic neuropathy (LHON) in Finland – a country that is among the best-studied genetic isolates in the world. During our long-term clinical follow-up period since 1970, we have so far identified 36 LHON families in Finland, comprised of almost 1000 family members. Counting the unaffected family members has been possible thanks to accessible genealogical records, and this has improved the accuracy of our penetrance figures by minimizing the sample bias. Our results, although confirming some well-known features of LHON, indicate that the overall penetrance of LHON is lower than previously estimated, and that affected females have a higher incidence of affected offspring compared to the unaffected females. The prevalence of LHON in Finland is 1:50 000, and one in 9000 Finns is a carrier of one of the three LHON primary mutations.
Similar content being viewed by others
Introduction
Leber hereditary optic neuropathy (LHON; OMIM1 # 5 35 000) is a maternally inherited eye disease that generally affects young adults with bilateral central vision loss leading to optic atrophy.2 The vast majority of LHON cases are associated with one of the three primary mutations in the mitochondrial DNA (mtDNA): MT-ND4/G11778A,3 MT-ND1/G3460A4, 5 or MT-ND6/T14484C.6 Prominent features in LHON pedigrees are incomplete penetrance and male bias among the affected individuals, reflecting the complex etiology of the disorder. Primary mutations are not sufficient in and of themselves to cause optic atrophy, but the additional factors – mutations in some nuclear gene(s) and/or epigenetic factors – that modify the risk of visual loss are still unknown.
We have performed an extensive epidemiological and penetrance study of LHON aiming to cover the entire population of Finland. We have combined thorough clinical expertise and a long-term follow-up of the Finnish LHON families over the period 1970–2004. As a country, Finland is well known for its particular suitability for this kind of study: it has a well-established population history and easily accessible and reliable genealogical records. Owing to a state-sponsored health-care system, medical records are also of high quality, and, in general, the patients and families have a favourable attitude towards health-care professionals and genetic research.
In this article, we describe the outcome of our epidemiological and penetrance study on the population of Finland.
Subjects and methods
Subjects
The clinical follow-up of the Finnish LHON families has been carried out by Turku University Central Hospital since 1970, conducted by one of the authors (E.N.). In our previous studies,7, 8 new LHON families have been diagnosed and the number of Finnish LHON families has increased to 36. This in all probability represents if not all, at least the overwhelming majority of the Finnish LHON families. Since the diagnosis of our first LHON patient in 1970 (by E.N.), Turku has been the leading clinical and molecular research centre for LHON in Finland. Ever since, we have maintained close contact networks between practising clinicians and researchers – an interaction that has guaranteed that even families with only a putative LHON diagnosis have been brought to our attention for further clinical and molecular genetic analyses.
We have also updated our previously identified pedigrees. By contacting the families and/or performing genealogical studies using Finnish parish records and civil registers, we have been able to add new individuals to them. In addition, the present clinical status of the family members has been ascertained. After this, we feel confident in concluding that any Finnish LHON cases that still may exist and our group is not aware of, are very few in number and most likely represent single cases, small families or distant branches of the previously diagnosed families.
The total number of subjects in the present study is 932 individuals (472 males and 460 females) after exclusion of all unaffected family members <5 years of age (ages were determined on 31 December 2004) or who had died still unaffected before reaching 5 years. In our series, 5 years is the earliest age at the time of diagnosis of LHON and, therefore, individuals younger than this were excluded. The oldest generations without reliable clinical diagnoses were also excluded.
The category of the affected is composed of cases with acute or subacute optic neuropathy with fundus and visual field findings suggestive of LHON as described previously.9, 10 Maternal relatives not assessed but reported as being unaffected were classified as unaffected. In LHON, affected individuals are usually aware of their ophthalmologic problems. Of the 932 individuals, 140 (108 males and 32 females) were affected with LHON, and of them, 108 (85 males and 23 females) were alive at the end of year 2004.
The unaffected individuals were divided into two age cohorts: one containing 792 individuals ≥5 years of age and the other containing 649 individuals ≥25 years of age. The other age limit, 25 years, can be considered as the average age of onset in LHON. This is based on the literature, as many authors for example,10, 11 have placed the mean age at onset between the ages of 22.0 years12 and 27.6 years.13
Mutation detection
After informed consent, venous blood samples for molecular genetic analyses were collected from the subjects during their regular follow-up visits to the clinic. The ethical committee of the Hospital District of Varsinais–Suomi approved the study. DNA was extracted from the blood with the standard methods (Nucleon BACC3 kit for blood and cell cultures, Amersham Biosciences, Little Chalfont, UK). The three primary mutations were tested by a standard restriction-fragment length polymorphism (RFLP) analysis of polymerase chain reaction (PCR) fragments. In homoplasmic pedigrees, maternal family members have been proven to carry the same mutation and therefore, not everyone needs to be tested.14 Thus, maternal relatives of an individual carrying a homoplasmic pathogenic mutation were concluded to be carriers of the same mutation. In the families in which the pathogenic mutation was unknown, the same rationale was used.
MtDNA sequencing
Complete mtDNA sequences were obtained from 35 probands; one proband was excluded because there was not enough DNA to perform the sequencing and no other family members were available. Each fragment was amplified in a standard 25-μl reaction. PCR products were purified for sequencing with the GFX™ PCR DNA and Gel Band Purification Kit according to the manufacturer's instructions (Amersham Biosciences, Little Chalfont, UK). The purified PCR products were sequenced. Sequence variants were searched by Sequencher v. 3.0 software (Gene Codes Corporation, Ann Arbor, MI, USA). Mitochondrial haplogroups were determined by RFLP as indicated,15, 16 and by hypervariable segments (HVS) I and II of the mtDNA (nt 16024–16383 and 57–372, respectively) sequencing.
Quantification of heteroplasmy
Heteroplasmy was determined by solid-phase minisequencing as described elsewhere.8 Individuals harbouring 1–99% of normal mtDNA were considered heteroplasmic and were included in the study. Individuals with <1% of mutant mtDNA were excluded. Where heteroplasmic individuals had family members with a homoplasmic mutation, these members were included with the heteroplasmic families.
Penetrance and prevalence analysis
We determined penetrance as the proportion of affected individuals from all maternally related family members. All pedigrees were evaluated separately. Additionally, combined penetrance values were calculated for the joint family material, for different primary mutations, for heteroplasmic families, for the two different age cohorts (≥5 and ≥25 years of age), and for males and females separately. Penetrances were also determined for males and females in each separate generation where there were ≥2 individuals of that particular gender. In addition, females with children were identified (unaffected offspring <25 years of age were excluded) and the number of each mother's affected and unaffected children was counted.
The minimum point prevalence values for LHON patients and primary mutation carriers in the Finnish population were calculated using the most recently published data in Finland – our total population was 5 236 611 individuals (49% males, 51% females) on 31 December 2004 (Statistics Finland).
Results
Distribution of primary mutations
The distribution of the primary mutations, homo- and heteroplasmy, and familial and sporadic pedigrees are shown in Table 1. Of the 36 Finnish LHON families, 24 (67%) harboured a primary mutation, distributed as follows: MT-ND4/G11778A 53% (19/36 families), MT-ND1/G3460A 11% (4/36) and MT-ND6/T14484C 3% (1/36). Among the 24 families with a primary mutation, the ND4/G11778A mutation represents 79% (19/24), the ND1/G3460A mutation 17% (4/24) and the ND6/T14484C mutation 4% (1/24) of the families. Out of these 24 families, heteroplasmy was detected in five of them (21%). The proportion of mutant mtDNA in individual members of heteroplasmic families varied from 2 to 98% among females and from 23 to 98% among males (A Puomila, unpublished data).8
In our series, we had 12 families (33%, 12/36) without a primary mutation. Owing to a confident clinical diagnosis of LHON in these families, they were also included in our study. In 11 families from which complete mtDNA sequences were obtained, none of the sequence variants fulfilled the criteria for a primary pathogenic LHON mutation (Table 2).
A relatively large number, a total of 15 families, were sporadic (42%, 15/36), for example with only one affected person diagnosed so far.
Only two of our 36 families are likely to be genealogically related to each other through a maternal ancestor, because they share an identical mtDNA control region sequence (Table 2). This suggests that the primary LHON mutations have arisen on several occasions in different mitochondrial backgrounds among the Finnish population.
Eight families (22%, 8/36) – six with ND4/G11778A, one with ND6/T14484C and one without a primary mutation – belong to mitochondrial haplogroup J (six to the subcluster J1 and two to J2). This makes the frequency of haplogroup J significantly higher among LHON patients than among the Finnish controls, among whom the frequency has been established as 6.3% (n=400)17 (χ2=12.0, P≤0.001).
Penetrance of optic atrophy
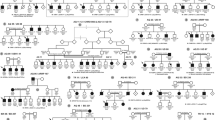
The penetrance figures for optic atrophy among the Finnish LHON families are shown in Table 3. In the first age cohort (≥5 years), the figures were as follows: in homoplasmic ND4/G11778A families, 26% for males and 6% for females, and in homoplasmic ND1/G3460A families, 19% for males and 9% for females. In the other age cohort (≥25 years), the penetrance figures were 31% for males and 8% for females in homoplasmic ND4/G11778A families, and 24% for males and 10% for females in homoplasmic ND1/G3460A families. Between separate families even with the same primary mutation, the penetrance is highly variable, as demonstrated in Figure 1. In the large family A, there is only one affected person, whereas in family B, several family members manifest optic atrophy. Family A belongs to haplogroup H that has been proposed to associate with low penetrance when occurring with the ND6/T14484C mutation.18 Family B represents haplogroup J which, on the other hand, has been linked to high penetrance in LHON.19, 20
Overall, the highest penetrance of optic atrophy was observed in the six ND4/G11778A-positive families belonging to mitochondrial haplogroup J: 37% of males (36/97) and 11% of females (10/95) at ≥5 years of age, and when only the age group of ≥25 years was considered, penetrances were even higher, 40% among males (36/90) and 12% among females (10/86).
To study whether the morbidity has changed over successive generations, penetrances were also determined for each generation separately in all informative pedigrees. No uniform trend towards increasing or decreasing penetrance in subsequent generations was observed in any of the families or different cohorts of families. Instead, among males, there were altogether 21 increasing, four stable, and 15 decreasing transitions from one generation to the next and among females, the corresponding figures were 7, 24 and 12. When data from both genders were combined, 28 of the penetrance figure transitions were increasing, 28 were stable and 27 were decreasing.
Male:female ratio
The well-known male preponderance in developing optic atrophy in LHON was evident. However, the ratio of affected males vs females was different depending on the mutation in question. Among homoplasmic ND4/G11778A-positive patients, the male:female ratio was 4.4:1 (80:18 individuals). The homoplasmic ND1/G3460A mutation was associated with a less prominent gender bias: the male:female ratio was 2.0:1 (10:5). In the ND6/T14484C family, there was only one affected male. Among individuals without a primary mutation, the ratio was 1.9:1 (17:9). The overall male:female ratio among all LHON patients was 3.4:1 (108:32).
Affected and unaffected females' risk of having affected offspring
To address whether the risk of affected offspring is greater among affected than unaffected females, we studied all mother–child pairs. We identified altogether 230 females who had offspring (unaffected offspring <25 years of age were excluded) (Table 4). Of the 230 mothers, 23 were affected and had a total of 52 descendants, that is on average 2.26 children each. The 207 unaffected mothers had 596 descendants (on average 2.88 each). Thus, unaffected mothers had slightly more children than affected mothers. Of the offspring of affected mothers, 38 (73%) were unaffected and 14 (27%) affected, whereas out of the offspring of unaffected mothers, 488 (82%) were unaffected and 108 (18%) affected. In the largest separate cohort, the homoplasmic ND4/G11778A-positive females, the affected mothers' ratio between unaffected and affected offspring was 28:10 (74–26%) whereas for the unaffected females the ratio was 312:75 (81–19%).
Group comparisons of the affected and unaffected children of the affected and unaffected mothers (carrying either ND4/G11778A, ND1/G3460A or no primary mutation) were performed using the χ2 test. The distribution was significant (P≤0.05) – affected mothers had a higher risk of having affected offspring than did unaffected mothers. It must be noted that the number of individuals remained <5 in three of the groups.
Prevalence
At the end of 2004, there were 108 LHON patients alive in our Finnish series. Thus, the minimum point prevalence of visual failure due to LHON within the Finnish population was 2.06 per 1 00 000 (95% confidence interval (CI) 1.80–2.40 per 1 00 000). At the same time, there were 560 living individuals carrying a LHON primary mutation in our series. This makes the minimum point prevalence of LHON primary mutations 10.69 per 1 00 000 among Finns (95% CI 10.10–11.30 per 1 00 000).
Discussion
When studying epidemiology, large and well-studied research populations are rare but extremely valuable. We have performed an entire-population-based survey on the epidemiology of LHON in Finland, a country that is among the best-studied genetic isolates in the world. We lack some inherited diseases such as cystic fibrosis or phenylketonuria that are common elsewhere, and have our own Finnish disease heritage that is due to our unique population history. However, none of these features are reflected in the molecular genetic parameters of LHON.
The frequencies of the LHON primary mutations in Finland do not strikingly differ from most of the other populations studied. In an extensive analysis of 159 LHON families from Northern Europe, the United Kingdom and Australia, 69% of the families carried the ND4/G11778A mutation, 13% the ND1/G3460A mutation and 14% the ND6/T14484C mutation21 compared with frequencies of 53, 11 and 3%, respectively, in our series. In Denmark, the ND6/T14484C mutation is as infrequent as in Finland (one family out of 30, whereas this frequency was one family out of 36 in our series).22
Prevalence
According to Chinnery et al,23 LHON is the most common mtDNA disorder. To date, data on the absolute prevalence of LHON in different populations are still sparse. In our 5.2-million population, LHON affects one in 48 487 individuals (2.06/1 00 000) and one in 9351 (10.69/1 00 000) carry a primary LHON mutation. Interestingly, this prevalence figure for LHON phenotype is remarkably close to the prevalence estimate of 1:50 000 based on clinical studies before the molecular genetic background of LHON was known.24 In a recent study of a population in North East England of 2.2 million,12 prevalence of visual failure due to LHON was one in 31 054 and of primary mutation carriers one in 8500. When the Finnish minimum point prevalence figures were compared with this data,12 the distribution of the primary mutation prevalence was not statistically significant (χ2=1.8, P=0.20). The minimum point prevalence of LHON phenotype, however, reached slight statistical significance between these two populations (χ2=8.6, P=0.044). In part, it could possibly indicate differences in, for example, the nuclear background between Finnish and English populations. The susceptibility alleles could be more common among the Englishmen. On the other hand, this difference may be due to restricting the English cohort to those at ‘working age’, that is ≤65 years of age. In Finland, individuals >65 years of age represent ∼16% of the population and therefore we included them in our survey. A remarkably lower penetrance for LHON (1:1 13 300) has been observed in Australia (a population of 17 million in 1994), but the prevalence for LHON primary mutations is of similar magnitude, one in 8500.14 All in all, at least in three Caucasoid populations, approximately 1 in 9000 people carry one of the three LHON primary mutations.
At the moment, there are approximately 80 000 visually handicapped people (according to the WHO (World Health Organization) criteria)25 in Finland.26 With the 108 LHON patients alive in Finland, it can be estimated that approximately one in every 740 visually handicapped Finnish individuals manifests LHON (0.14%). In Australia, LHON accounts for 0.42–2% of invalid blind people.11
Penetrance
In an adult-onset disorder, penetrance values may be highly variable depending on the age limits used in the studies. This is clearly demonstrated in our study, where two different age cohorts result in different penetrance figures. However, we consider excluding young unaffected individuals (<25 years of age) to be reasonable because it can be assumed that a number of them will be affected when reaching the age of 25. The most comparable penetrance values are therefore obtained within one study that is based on a sufficient quantity of research material. In our series, there were substantial penetrance differences even among families carrying the same mutation. This has been observed in other populations as well.27
Penetrance in specific cohorts
In our study, the penetrance of optic atrophy was lower than in earlier studies. Among all our families, the penetrance was 27% in males and 8% in females, whereas the majority of earlier reports have given penetrances as high as ∼50–60% in males and ∼10–20% in females.22, 28, 29, 30, 31. Our previous estimation was 45% for males and 18% for females.32 The present lower penetrances are probably due to the fact that in this study, we have done our best to trace all maternal members of the families, which has increased the number of unaffected relatives included. The lowest overall penetrances have been reported in a genealogical study by Mackey: 20% for males and 4% for females.14
When the penetrance values were calculated for each generation in all informative pedigrees, we found no evidence that the risk of developing LHON systematically changes over successive generations. In our series of 83 transitions from a previous generation to the subsequent in 25 informative pedigrees, the penetrance figures fluctuated seemingly randomly. No evident consistent changes towards either an increasing or decreasing risk of optic atrophy were seen in all the pedigrees combined or in any given cohort. This is in opposition to the results of Mackey14 and Sadun et al33 who have shown cases where penetrance decreases in younger generations of the pedigrees. However, in the study of Sadun et al33, there was only one, albeit very large, pedigree with nearly 300 family members. In a case with just one family, the effect of specific nuclear and/or environmental factors cannot be ruled out.
LHON families without a primary mutation In our series, the penetrance values in families without a primary mutation were quite similar to those with a primary mutation. Families with no known LHON primary mutation but still having a confident clinical diagnosis of LHON continue to be enigmas. We have 12 such cases, and recently Fauser et al34 reported 14 LHON patients with typical clinical features of LHON but no primary mutation. Among our 12 cases, six are sporadic and six are familial. In four of these families, there is a maternal inheritance pattern (e.g. in one family, there are seven affected maternal relatives) and two are male sib-pairs. Mackey14, too, has reported two male sib-pairs, neither carrying any of the primary mutations. In 11 of our families, none of the mtDNA sequence variants fulfilled the criteria for a primary pathogenic mtDNA mutation (Table 2). One intriguing and also diagnostically challenging possibility that remains is that there is some bilateral optic atrophy; clinically very similar to LHON but not associated with mtDNA defects. Thus the underlying genetic defect may be nuclear and inheritance simply mimics maternal inheritance.
Sporadic cases and heteroplasmic families Despite LHON being an inherited disease, many diagnosed individuals represent isolated cases even in large families. The proportion of sporadic cases has been reported to be 57% for the ND4/G11778A, 22% for the ND1/G3460A and 35% for the ND6/T14484C mutation.13, 35 In our series, the proportion of sporadic families is 15 out of 36 families (42%). The large degree of sporadicity, however, is not exceptional and restricted solely to Finns. In two previous studies, similar results were obtained: 36%30 and 40%31, although this is in contrast with 2% in Australia.14
Sporadic cases are often heteroplasmic, and correspondingly, four out of our five heteroplasmic families are sporadic. Heteroplasmy has been proposed to be a penetrance-decreasing factor,36 although individuals with a relatively low proportion of mutant mtDNA in their blood can still become affected and females with extremely low proportion of mutant mtDNA can have affected offspring.37 In the present survey, the heteroplasmic families were too small to provide any comparable penetrance figures.
Haplogroup J families Out of the Finnish LHON families, eight (22%; six ND4/G11778A-positive, one ND6/T14484C-positive and one without a primary mutation) belong to mitochondrial haplogroup J, making haplogroup J significantly more frequent among our LHON families than among the Finnish controls (6.3% in Niemi et al17). This, and especially the high frequency of ND4/G11778A- and ND6/T14484C-positive families in haplogroup J, has been seen in previous studies as well.7, 19, 20, 38 In our series, the highest penetrance was seen in the ND4/G11778A mutation which occurred in the haplogroup J background, in accordance with previous studies.19, 20
Recently, several studies on neurological diseases have supported the view that mtDNA variation in certain haplogroups may not be completely neutral but may either increase or decrease the risk of developing the symptoms.39, 40 Reynier et al41, for example, suggest that haplogroup J could be a genetic factor for visual impairment. Despite these haplogroup J-associated findings in LHON, the observed phenomenon is difficult to explain in the light of the present limited knowledge of the role which mtDNA variation ultimately plays. The hypothesis is that certain mtDNA polymorphisms are, by themselves, neutral but in specific combinations together with pathogenic mtDNA mutations they might be deleterious and increase the risk of disease expression or produce a more severe clinical phenotype.42
Male preponderance
The well-established concept of male preponderance in LHON was also seen in this study. The male:female ratios of the affected family members are listed in Table 5. The Finnish ratios are in accordance with those obtained earlier from other Caucasoid populations. For example, our ratios were compared with the English material12 and the distributions were not statistically significantly different for any of the groups – ND4/G11778A-positive, ND1/G3460A-positive, all primary mutations combined or the whole material combined (data not shown).
When the primary mutations are compared, the male preponderance is the most prominent in patients with the ND6/T14484C mutation and less obvious in patients with the ND1/G3460A mutation (Table 5). This may be related to the differences in the biochemical consequences of the primary mutations. In an extensive respiration and enzymological analysis on lymphoblast and cybrid mitochondria isolated from 19 LHON patients,46 the most severe biochemical defect was detected for the ND1/G3460A mutation, an intermediate one for the ND4/G11778A mutation, and the mildest for the ND6/T14484C mutation. Female carriers of the biochemically most severe ND1/G3460A mutation seem to become affected almost as often as males, whereas among carriers of other, less severe mutations, a greater proportion of females remains unaffected. Man et al38 have proposed that the ND1/G3460A is a ‘stronger’ mutation and less susceptible to the epistatic and epigenetic factors influencing the expression of the ND4/G11778A and ND6/T14484C mutations.
Affected and unaffected females' risk of having affected offspring
In our family material, there were 230 females with offspring (unaffected offspring <25years of age were excluded). The unaffected mothers had slightly more descendants than the affected mothers. The affected females had a higher incidence of affected offspring than the unaffected females (P=0.034). This is in accordance with the study of Harding et al30 in which 58 females were analysed and affected mothers had statistically significantly more affected offspring than did unaffected mothers.
Conclusions
Our survey was performed to obtain a clear and defined picture of the epidemiology, prevalence and penetrance of LHON in a well-studied population of an entire country. Adult-onset mitochondrial disorders, such as LHON, are a problematic field in epidemiology, but in a well-known and accessible study population, variability due to sampling effects can be avoided and accuracy is therefore improved. Complete understanding of these diseases can be achieved only when the interplay between our two genomes, nuclear and mitochondrial, in each disorder has been clarified.
References
OMIM™: Online Mendelian inheritance in man. McKusic-Nathans Institute for Genetic Medicine, Johns Hopkins University (Baltimore, USA); and National Center for Biotechnology Information, National Library of Medicine (Bethesda, USA). Available at http://www.ncbi.nlm.nih.gov/omim/.
Nikoskelainen EK : Clinical picture of LHON. Clin Neurosci 1994; 2: 115–120.
Wallace DC, Singh G, Lott MT et al: Mitochondrial DNA mutation associated with Leber's hereditary optic neuropathy. Science 1988; 242: 1427–1430.
Howell N, Bindoff LA, McCullough DA et al: Leber hereditary optic neuropathy: identification of the same mitochondrial ND1 mutation in six pedigrees. Am J Hum Genet 1991; 49: 939–950.
Huoponen K, Vilkki J, Aula P, Nikoskelainen EK, Savontaus M-L : A new mtDNA mutation associated with Leber's hereditary optic neuroretinopathy. Am J Hum Genet 1991; 48: 1147–1153.
Johns DR, Neufeld MJ, Park RD : An ND-6 mitochondrial DNA mutation associated with Leber hereditary optic neuropathy. Biochem Biophys Res Commun 1992; 18: 1551–1557.
Lamminen T, Huoponen K, Sistonen P et al: mtDNA haplotype analysis in Finnish families with Leber hereditary optic neuropathy. Eur J Hum Genet 1997; 5: 271–279.
Puomila A, Viitanen T, Savontaus ML, Nikoskelainen E, Huoponen K : Segregation of the ND4/11778 and the ND1/3460 mutations in four heteroplasmic LHON families. J Neurol Sci 2002; 205: 41–45.
Nikoskelainen E, Hoyt WF, Nummelin K : Ophthalmoscopic findings in Leber's hereditary optic neuropathy. II. The fundus findings in the affected family members. Arch Ophthalmol 1983; 101: 1059–1068.
Nikoskelainen EK, Huoponen K, Juvonen V, Lamminen T, Nummelin K, Savontaus M-L : Ophthalmologic findings in Leber hereditary optic neuropathy, with special reference to mtDNA mutations. Ophthalmology 1996; 103: 504–514.
Mackey DA, Buttery RG : Leber hereditary optic neuropathy in Australia. Aust N Z J Ophthalmol 1992; 20: 177–184.
Man PYW, Griffiths PG, Brown DT, Howell N, Turnbull DM, Chinnery PF : The epidemiology of Leber hereditary optic neuropathy in the North East of England. Am J Hum Genet 2003; 72: 333–339.
Newman NJ, Lott MT, Wallace DC : The clinical characteristics of pedigrees of Leber's hereditary optic neuropathy with the 11778 mutation. Am J Ophthalmol 1991; 111: 750–762.
Mackey DA : Epidemiology of Leber's hereditary optic neuropathy in Australia. Clin Neurosci 1994; 2: 162–164.
Torroni A, Huoponen K, Francalacci P et al: Classification of European mtDNAs from an analysis of three European populations. Genetics 1996; 144: 1835–1850.
Macaulay V, Richards M, Hickey E et al: The emerging tree of West Eurasian mtDNAs: a synthesis of control-region sequences and RFLPs. Am J Hum Genet 1999; 64: 232–249.
Niemi AK, Hervonen A, Hurme M, Karhunen PJ, Jylhä M, Majamaa K : Mitochondrial DNA polymorphisms associated with longevity in a Finnish population. Hum Genet 2003; 112: 29–33.
Howell N, Herrnstadt C, Shults C, Mackey DA : Low penetrance of the 14484 mutation when it arises in a non-haplogroup J mtDNA background. Am J Med Genet A 2003; 119A: 147–151.
Brown MD, Sun F, Wallace DC : Clustering of Caucasian Leber hereditary optic neuropathy patients containing the 11778 or 14484 mutations on an mtDNA lineage. Am J Hum Genet 1997; 60: 381–387.
Torroni A, Petrozzi M, D'Urbano L et al: Haplotype and phylogenetic analyses suggest that one European-specific mtDNA background plays a role in the expression of Leber hereditary optic neuropathy by increasing the penetrance of the primary mutations 11778 and 14484. Am J Hum Genet 1997; 60: 1107–1121.
Mackey DA, Oostra RJ, Rosenberg T et al: Primary pathogenic mtDNA mutations in multigeneration pedigrees with Leber hereditary optic neuropathy. Am J Hum Genet 1996; 59: 481–485.
Bech N, Rosenberg T : Leber's hereditære opticus neuropathi. Ophthalmolog 1998; 2: 3–4.
Chinnery PF, Johnson MA, Wardell TM et al: The epidemiology of pathogenic mitochondrial DNA mutations. Ann Neurol 2000; 48: 188–193.
Orphanet. website on rare diseases and orphan drugs. See http://www.orpha.net/data/patho/GB/uk-LHON.pdf.
WHO: International Classification of Impairments, Disabilities, and Handicaps. Geneva: World Health Organization, 1980, chap. 5.
Ojamo M : Näkövammarekisterin vuosikirja. Stakes, Näkövammaisten keskusliitto ry 2003.
Howell N, Kubacka I, Halvorson S, Mackey D : Leber's hereditary optic neuropathy: the etiological role of a mutation in the mitochondrial cytochrome b gene. Genetics 1993; 140: 285–302.
van Senus AHC : Leber's disease in the Netherlands. Doc Ophthalmol 1963; 17: 1–162.
Seedorff T : The inheritance of Leber's disease. A genealogical follow-up study. Acta Ophthalmol (Copenh) 1985; 63: 135–145.
Harding AE, Sweeney MG, Govan GG, Riordan-Eva P : Pedigree analysis in Leber hereditary optic neuropathy families with a pathogenic mtDNA mutation. Am J Hum Genet 1995; 57: 77–86.
Man PY, Turnbull BM, Chinnery PF : Leber hereditary optic neuropathy. J Med Genet 2002; 39: 162–169.
Nikoskelainen EK, Savontaus ML, Wanne OP, Katila MJ, Nummelin KU : Leber's hereditary optic neuroretinopathy, a maternally inherited disease. A genealogic study in four pedigrees. Arch Ophthalmol 1987; 105: 665–671.
Sadun AA, Carelli V, Salomao SR et al: Extensive investigation of a large Brazilian pedigree of 11778/haplogroup J Leber hereditary optic neuropathy. Am J Ophthalmol 2003; 136: 231–238.
Fauser S, Luberichs J, Besch D, Leo-Kottler B : Sequence analysis of the complete mitochondrial genome in patients with Leber's hereditary optic neuropathy lacking the three most common pathogenic DNA mutations. Biochem Biophys Res Commun 2002; 295: 342–347.
Johns DR, Katrinka LH, Miller NR, Smith KH : Leber's hereditary optic neuropathy: clinical manifestations of the 14484 mutation. Arch Ophthalmol 1993; 111: 495–498.
Chinnery PF, Andrews RM, Turnbull DM, Howell N : Leber hereditary optic neuropathy: does heteroplasmy influence the inheritance and expression of the G11778A mitochondrial DNA mutation? Am J Med Genet 2001; 98: 235–243.
Huoponen K, Puomila A, Savontaus M-L, Mustonen E, Kronqvist E, Nikoskelainen E : Genetic counseling in Leber hereditary optic neuropathy (LHON). Acta Ophthalmol Scand 2002; 80: 38–43.
Man PYW, Howell N, Mackey DA et al: Mitochondrial DNA haplogroup distribution within Leber hereditary optic neuropathy pedigrees. J Med Genet 2004; 41: e41.
Giacchetti M, Monticelli A, De Biase I et al: Mitochondrial DNA haplogroups influence the Friedreich's ataxia phenotype. J Med Genet 2004; 41: 293–295.
Autere J, Moilanen JS, Finnilä S et al: Mitochondrial DNA polymorphisms as risk factors for Parkinson's disease and Parkinson's disease dementia. Hum Genet 2004; 115: 29–35.
Reynier P, Penisson-Besnier I, Moreau C et al: MtDNA haplogroup J: a contributing factor of optic neuritis. Eur J Hum Genet 1999; 7: 404–406.
Chinnery PF, Howell N, Andrews RM, Turnbull DM : Mitochondrial DNA analysis: polymorphisms and pathogenicity. J Med Genet 1999; 36: 505–510.
Black GC, Craig IW, Oostra RJ et al: Leber's hereditary optic neuropathy: implications of the sex ratio for linkage studies in families with the 3460 ND1 mutation. Eye 1995; 9: 513–516.
MacMillan C, Kirkham T, Fu K et al: Pedigree analysis of French Canadian families with T14484C Leber's hereditary optic neuropathy. Neurology 1998; 50: 417–422.
Riordan-Eva P, Sanders MD, Govan GG, Sweeney MG, Da Costa J, Harding AE : The clinical features of Leber's hereditary optic neuropathy defined by the presence of a pathogenic mitochondrial DNA mutation. Brain 1995; 118: 319–337.
Brown MD, Trounce IA, Jun AS, Allen JC, Wallace DC : Functional analysis of lymphoblast and cybrid mitochondria containing the 3460, 11778, or 14484 Leber's hereditary optic neuropathy mitochondrial DNA mutation. J Biol Chem 2000; 275: 39831–39836.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Puomila, A., Hämäläinen, P., Kivioja, S. et al. Epidemiology and penetrance of Leber hereditary optic neuropathy in Finland. Eur J Hum Genet 15, 1079–1089 (2007). https://doi.org/10.1038/sj.ejhg.5201828
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201828
Keywords
This article is cited by
-
Mitochondrial heteroplasmic shifts reveal a positive selection of breast cancer
Journal of Translational Medicine (2023)
-
Magnetic resonance imaging negative myelopathy in Leber’s hereditary optic neuropathy: a case report
BMC Neurology (2022)
-
Natural history of patients with Leber hereditary optic neuropathy—results from the REALITY study
Eye (2022)
-
Leber hereditary optic neuropathy—new insights and old challenges
Graefe's Archive for Clinical and Experimental Ophthalmology (2021)
-
Safety of Intravitreal Gene Therapy for Treatment of Subjects with Leber Hereditary Optic Neuropathy due to Mutations in the Mitochondrial ND4 Gene: The REVEAL Study
BioDrugs (2021)